Vinco-T4
Phage size does Matter…
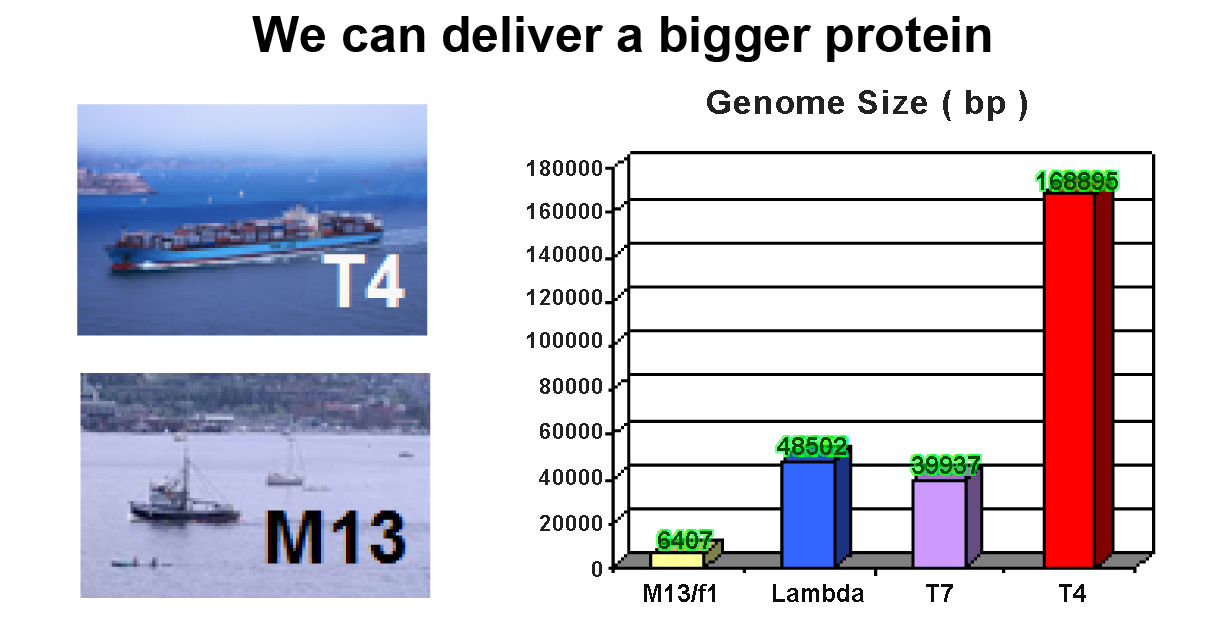
It represents the maximum potential capacity of the insertion length (bp). Small phage particle cannot afford the over-size DNA cloning. 
 |
| It is critical for cloning large and multi-mers ( e.g. VEGFR, |
| Flu vaccine M2e homotetramers ). |
| Small phage (e.g. M13, T7 ) cannot afford the over-size DNA cloning. |

Phage display is a powerful technique for identifying peptides or proteins that have desirable binding properties. T4 phage display system will make an impact in various phage display systems for reasons of: Phage T4 display system makes it possible to construct macromolecular complexes that can simultaneously display two proteins or peptides with different lengths and properties in hundreds to thousands of identical copies of designated protein antigens, domains, or enzymatic activities on a single macromolecular particle. It is anticipated that such complexes will find widespread applications in vaccine development, protein-protein binding interactions, biotechnology and in drug discovery research. The powerful research values are especially included in protein /antibody libraries construction, that greatly facilitate protein / antibodies optimization, identification of structurally constrained peptide, target protein and its gene discovery, select cytokine receptor agonists, and for drug receptor agonists / inhibitors ligand binding site / active domain identification. In biotechnology, assembling massively multimeric copies of rare, expensive, or unstable antibody / enzymatic activities onto these macro-molecular platforms should make it possible to generate novel biotechnological reagents and techniques [12]. In recent years, the use of surface-display vectors for displaying polypeptides on the surface of bacteriophage, combined with in vitro selection technologies, has transformed the way in which we generate and manipulate protein domains such as antigens, antibodies and enzymes [30]. Phage T4 display is based on expressing recombinant proteins or peptides fused to T4 phage coat dispensable protein Hoc and Soc. the genetic information encoding for the displayed molecule is physically linked to its product via the displaying particle. Using this technology, we are now able to design repertoires of ligands from scratch and use the power of affinity selection to select those protein domains having the desired biological properties from a large excess of irrelevant T4 phage particles. With T4 phage display, customer-made proteins (fused antigen peptides, antibodies, enzymes, DNA-binding proteins) may be synthesized and selected to acquire the desired affinity of binding and specificity for in vitro and in vivo diagnosis, various antigenic determinants, heterologous enzymes, single-chain antibodies, and combinatorial peptide libraries, in the future for immunotherapy of human disease. This explains the basis of T4 phage bipartite display and investigates of the new phenomenon compared with plasmid, Ff phage expression and display. T4 phage two sites synchronously display technique is prospecting to make contributions to biotechnological applications. This technique focuses mainly on T4-antibody library constructions and screening. The approach involves the simultaneous selection of VL and VH domains, respectively, where T4 phage display would have the greatest impact on antibody engineering, and vaccine development.
Recombinant antibodies have been displayed on bacteriophage [14], lytic phage [15], bacteria [16], yeast [17], ribosomes[18] provide invaluable data for both basic research and medical diagnostics. Antibodies have opened up new possibilities for high-throughput profiling of protein expression. The past years have seen a number of advances in library design [31]. Phage–antibody libraries have so far yielded antibodies to several hundred targets, including cell-surface markers, peptide hormones, other human
T4 Phage display system (Successful Antigens, Antibodies, and enzymes):
- HIV: V3
- Polio: VP1 (320 aa/960 bp)
- FMDV: P1 (839 aa/2517 bp)
- CSFV: E2 (374 aa/1122 bp)
- IBDV
- AIV
- NDV
- EWL-IgG
- VH-link-VL
- T4 DNA Ligase
- FMDV 3C antigen-proteinase
- HIV: receptor domain 1+2